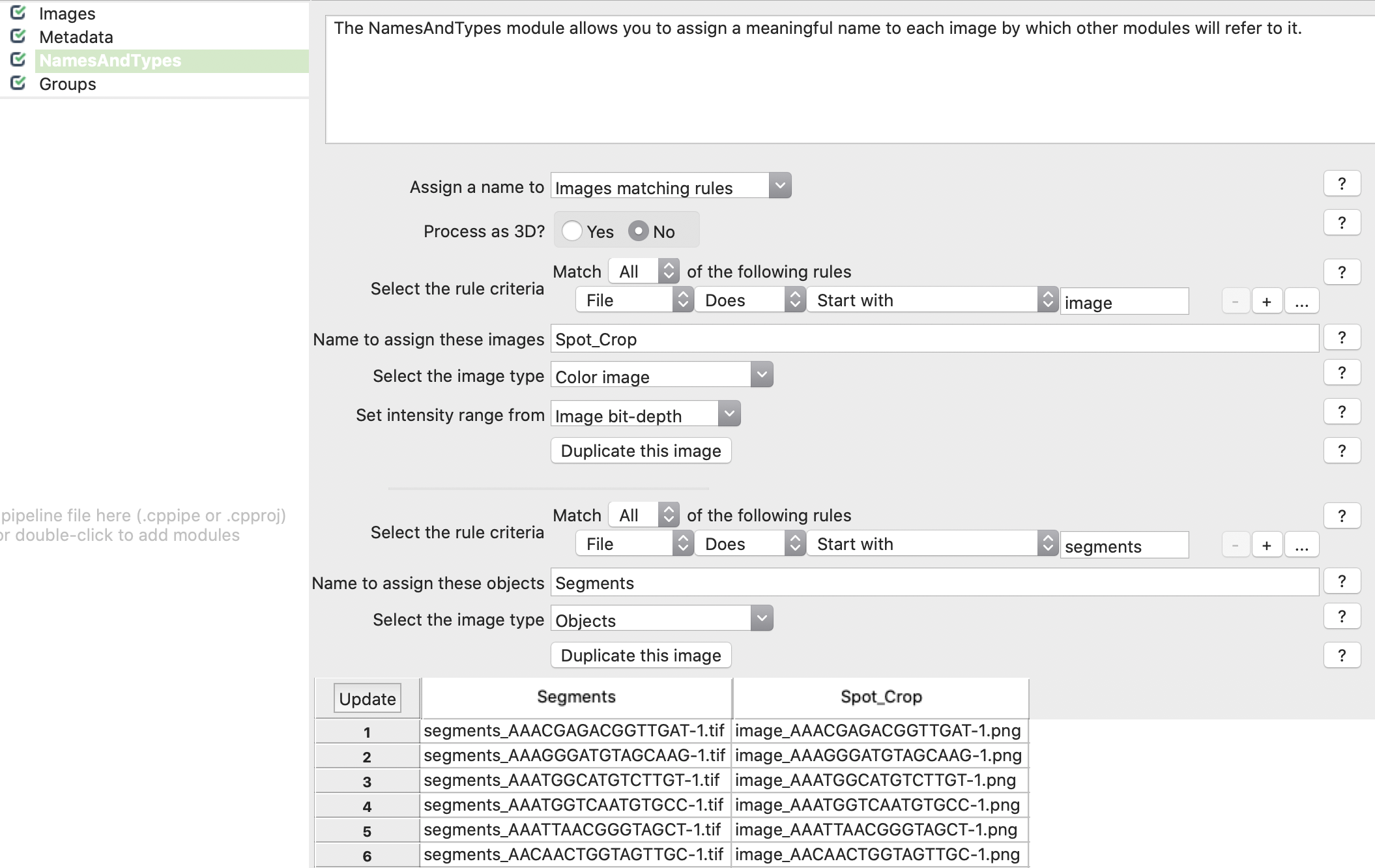
Plate Viewer displays data according to the. Classifier enables cell and field-of-view-level classification of multiple phenotypes using popular supervised machine learning models. I wanted to store the CSV files locally on the computer rather than using the mySQL database. So I tried using the ExportToDatabase function for the first time at the end of my pipeline. It was originally published for the visualization of phenotypic kidney features in zebrafish, but the workflow is generic by design and can be reused for any quantitative feature set. CellProfiler Analyst’s primary tools: Image Gallery displays (full and cell) images with a variety of filter options and can be used interactively with other tools. Click on the magnifying glass to see how Cellprofiler reads your data. I’m try to create a properties file to use for CPA. The workflow is interactive and so selecting in one panel of the figure will highlight in the other panel too. The % is shown in the title of the PCA plot. Here, we provide links to pipelines used in some published studies, as well as the version of CellProfiler used to create them. Otherwise the PCA plot will not be representative of the original data-distribution. CellProfiler enables reproducible research because a saved pipeline includes all the modules and settings. One option is to normalize the values (min/max or Z-score).Īlso make sure that the resulting PCA represents a decent % of the original data variance (at least 70%). If you plan to make these files accessible via the web, you can enter a url prefix here. The original features should cover similar value range, to make sure the PCA is not biased towards the large values features. KNIME workflow to visualize a dataset described by multiple quantitative features (ex: a list of samples or cells, each described with multiple morphological features) as a 3D cloud of points (each point corresponding to one sample/cell) as well as a line plot (1 line per sample/cell).įor the 3D plot, the workflow uses Principal Component Analysis (PCA) for dimensionality reduction, ie it simplifies the information for each sample from n-features to 3 pseudo-features which are used as x,y,z-coordinates for each sample. Export analytical data to text or CSV files.Select an extensive range of measurement parameters.Analyse areas via data thresholding using amount of x-ray counts, phase maps, height, or other material properties.Measure and quantify data across multiple layers.Generate profile (cross section) views of multimodal data.Customisation of colour palettes, data overlays, image rendering options, and document display.3D visualisation of topography combined with AFM material properties, EM images, and EDS & EBSD map overlays.Correlate both sets of data using intuitive image overlays and image matching tools.Import data from AZtec using the H5oina file format.Interactive display of multi-layer correlated dataĪnalytical tools for metadata interrogation It provides the tools you need to correlate data from different microscopes, visualise multi-layered data in 2D and 3D, and conduct correlative analyses.Ĭombining data from different imaging modalities (e.g. Relate is a correlative software package optimised to work with EM, EDS, EBSD, & AFM data and images.
